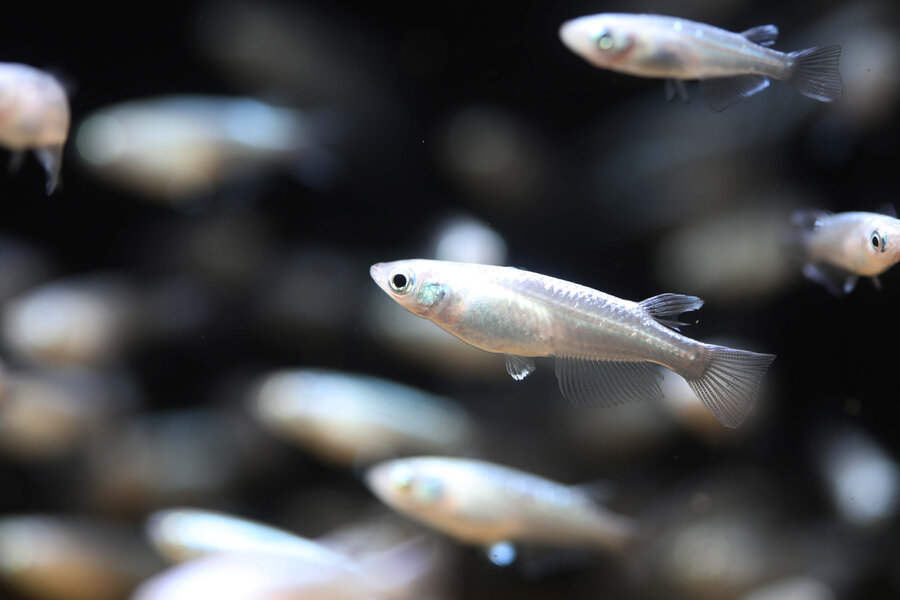
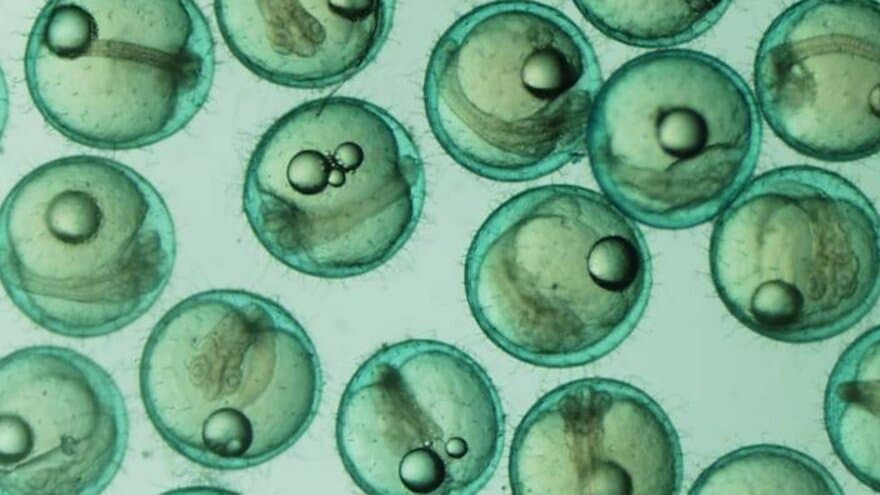
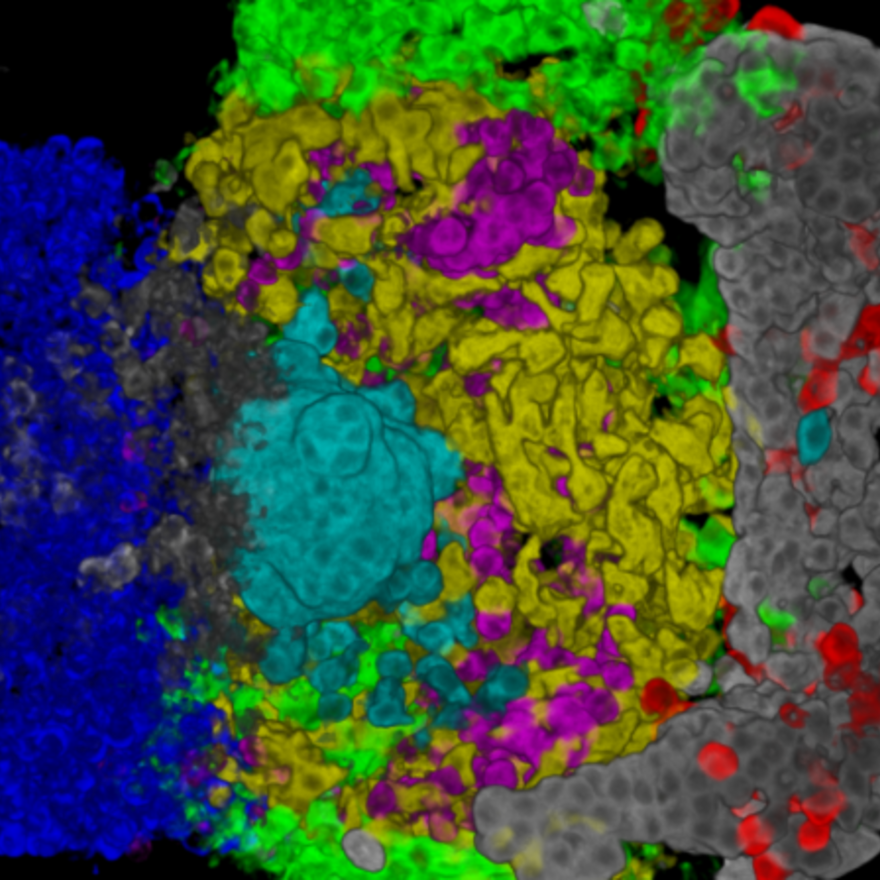
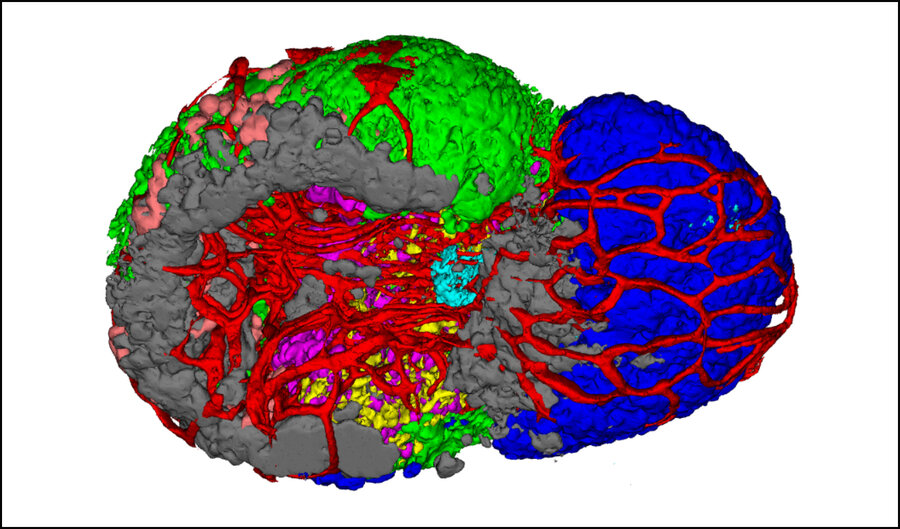
This project aims to develop a highly detailed description of the pituitary gland of the medaka fish.
Our main objective
This project provides an overview of the mRNA content of each pituitary cell types in the male and female adult medaka.
It also provides a detail 3D map of the distribution of the different endocrine cell populations in the juvenile and adult medaka.
About the Medaka Pituitary Atlas Project
3D spatial distribution
Tutorials - how to use the 3D model
Single Cell Transcriptome
Participants
Publications
2021
3D Atlas of the Pituitary Gland of the Model Fish Medaka (Oryzias latipes)
Frontiers in Endocrinology
2021
2020
Labeling of blood vessels in the teleost brain and pituitary using cardiac perfusion with a DiI-fixative
JoVE (Journal of Visualized Experiments), e59768
2020