A main research focus of LMG is bacteriocins, antimicrobial peptides produced by bacteria. Our work covers all aspects of bacteriocins research, from screening, purification, identification and characterization.
We aim to understand how bacteriocins interact with their molecular targets (receptors) on bacterial cells and how these interactions results in cell death and growth inhibition. Improved knowledge on the mechanism and targets of bacteriocins is important for developing these peptides for practical applications such as food preservation or as novel drugs.
- We have a high-throughput screening technique for bacteriocins targeting pathogens of interest
- Our major pathogens of focus are MRSA, VRE and Listeria
- We have a well-established platform for bacteriocin studies covering both biochemical and genetic aspects
- We have established skin infection models in mice for therapeutic studies
- We have diverse luciferase-tagged pathogens and an IVIS imaging camera for studying infection development in animal models
Another major focus of LMG is Staphyloccus aureus: cell division proteins, essential proteins, stress-response systems and bacteriocin targets. S. aureus is a major opportunistic pathogen in humans and antibiotic resistance is an increasing problem with strains such as methicillin-resistant S. aureus (MRSA) or vancomycin-resistance S. aureus (VRSA). Consequently, there is an urgent need for new antimicrobials against this species. A better understanding of essential genes and their functions in S. aureus will be cruciual for finding new and novel antimicrobial targets.
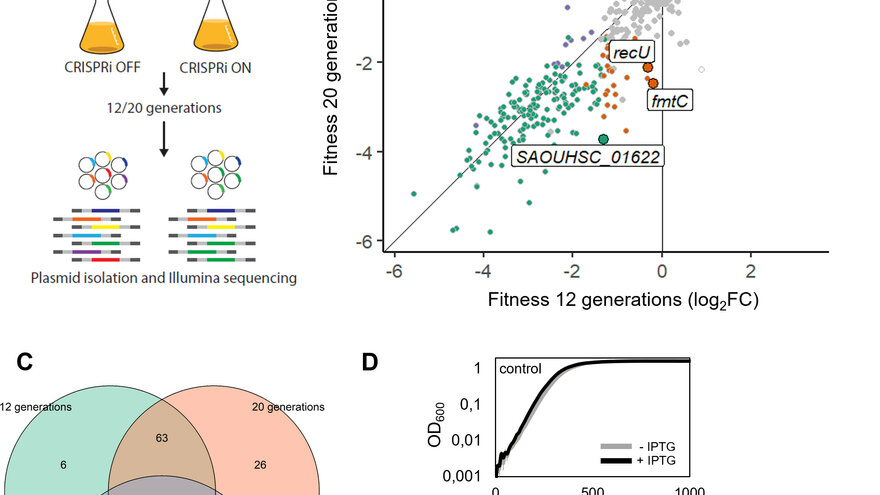
- We have established a genome-wide CRISPRi screen which allows us to study the essentiality of genes under different conditions
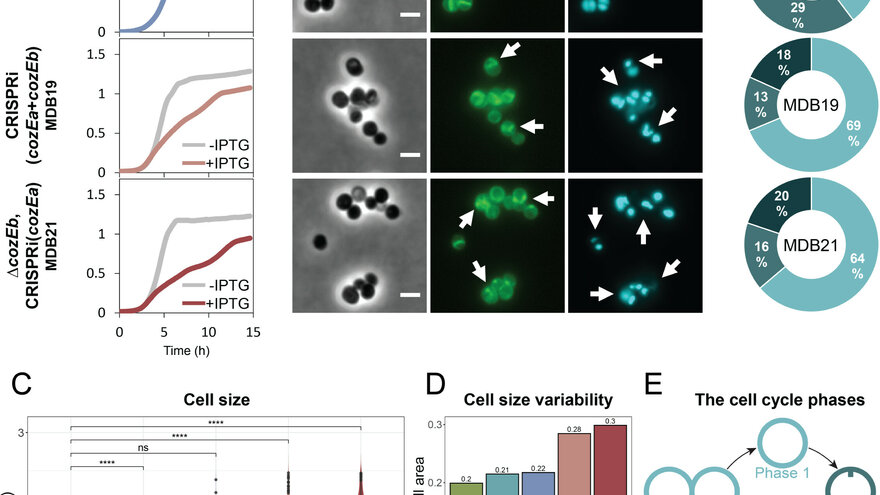
- We have modern fluorescence and confocal microscopes to study localization and cell morphology
- Tools for efficient genetic manipulation and gene expression in S. aureus
Recent publications
About LMG
News
Bruker bakterienes egne våpen mot resistente bakterier

Photo: Tonje Halvorsen Walde/NMBU Bakterienes egne våpen kan brukes mot andre bakterier som gjør oss syke
Photo: Alexander Benjaminsen/NMBU
Publications
List of publications:
Liu X., deBakker V., Heggenhougen MV., Mårli MT., Frøynes AH., Salehian Z., Porcellato D., Morales Angeles D., Veening J., Kjos M. Genome-wide CRISPRi screens for high-throughput fitness quantification and identification of determinants for dalbavancin susceptibility in Staphylococcus aureus. mSystems 0:e01289-23.https://doi.org/10.1128/msystems.01289-23
Audshasai, T., Coles, J. A., Panagiotou, S., Khandaker, S., Scales, H. E., Kjos, M., Baltazar, M., Vignau, J., Brewer, J. M., Kadioglu, A., Yang, M. 2022. Streptococcus pneumoniae Rapidly Translocate from the Nasopharynx through the Cribriform Plate to Invade the Outer Meninges. mBio 13:e01024-22.https://doi.org/10.1128/mbio.01024-22
Cacace, E., Kim, V., Varik, V. et al. Systematic analysis of drug combinations against Gram-positive bacteria. Nat Microbiol 8, 2196–2212 (2023). https://doi.org/10.1038/s41564-023-01486-9
Arbulu, S., Kjos, M. Revisiting the Multifaceted Roles of Bacteriocins. Microb Ecol 87, 41 (2024). https://doi.org/10.1007/s00248-024-02357-4
Nielsen TK, Petersen IB, Xu L, Barbuti MD, Mebus V, Justh A, Alqarzaee AA, Jacques N, Oury C, Thomas V, Kjos M, Henriksen C, Frees D. 2024. The Spx stress regulator confers high-level β-lactam resistance and decreases susceptibility to last-line antibiotics in methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 68:e00335-24.https://doi.org/10.1128/aac.00335-24
Reinseth I., Diep DB, Kjos M., Tønnesen H. H., Carlsen H., Exploring the feasibility of bacteriocins EntK1 and EntEJ97s in treatment of systemic vancomycin resistant enterococci infections in mice, Journal of Applied Microbiology, Volume 135, Issue 3, March 2024, lxae054, https://doi.org/10.1093/jambio/lxae054
Oftedal, T.F., Diep, D.B. & Kjos, M. Design of Novel Saposin-like Bacteriocins Using a Hybrid Approach. Probiotics & Antimicro. Prot. (2024). https://doi.org/10.1007/s12602-024-10264-w
Barbuti MD, Lambert E, Myrbråten IS, Ducret A, Stamsås GA, Wilhelm L, Liu X, Salehian Z, Veening J, Straume D, Grangeasse C, Perez C, Kjos M. 0. The function of CozE proteins is linked to lipoteichoic acid biosynthesis in Staphylococcus aureus. mBio 0:e01157-24.https://doi.org/10.1128/mbio.01157-24
Wolden R, Ovchinnikov KV, Venter HJ, Oftedal TF, Diep DB, Cavanagh JP (2023). The novel bacteriocin romsacin from Staphylococcus haemolyticus inhibits Gram-positive WHO priority pathogens. Microbiol Spectr. 2023 Oct 31:e0086923. doi: 10.1128/spectrum.00869-23.
Cacace E, Kim V, Knopp M, Tietgen M, Brauer-Nikonow A, Inecik K, Mateus A, Milanese A, Mårli MT, Mitosch K, Selkrig J, Brochado AR, Kuipers O, Kjos M, Zeller G, Savitski M, Göttig S, Huber W, Typas A (2023) High-throughput profiling of drug interactions in Gram-positive bacteria. Preprint: https://www.biorxiv.org/content/10.1101/2022.12.23.521747v1.
Hauge IH, Sandegren V, Ruud Winther A, Bøe CA, Salehian Z, Håvarstein LS, Kjos M, Straume D (2023) A novel proteinaceous molecule produced by Lysinibacillus sp. OF-1 depends on the Ami oligopeptide transporter to kill Streptococcus pneumoniae. Microbiology. 169(3):001313. https://doi.org/10.1099/mic.0.001313
Disen Barbuti M, Myrbråten I, Morales Angeles D, Kjos M (2022) An updated overview of the cell cycle promesses in Staphylococcus aureus. MicrobiologyOpen. In press. doi.org/10.1002/mbo3.1338
Gao Y, Kjos M, Arntzen MØ, Bakken LR, Frostegård Å (2023) Denitrification by bradyrhizobia under feast and famine and the role of the bc1 complex in securing electrons for N2O reduction. Appl Environ Microbiol. https://doi.org/10.1128/aem.01745-22. Preprint : https://www.biorxiv.org/content/10.1101/2022.09.29.510233v1
Audshasai T, Coles JA, Panagiotou S, Khandaker S, Scales HE, Kjos M, Baltazar M, Vignau, Brewer JM, Kadioglu A, Yang M (2022). Streptococcus pneumoniae rapidly translocates from the nasopharynx through the cribriform plate to invade and inflame the dura. mBio. 3(4):e0102422. doi.org/10.1128/mbio.01024-22
Myrbråten I, Stamsås GA, Chan H, Morales Angeles D, Knutsen TM, Salehian Z, Shapaval O, Straume D, Kjos M. (2022) SmdA is a novel cell morphology determinant in Staphylococcus aureus. mBio. doi.org/10.1128/mbio.03404-21. Commentary: doi.org/10.1128/mbio.00737-22
Gallay C, Sanselicio S, Anderson ME, Soh YM, Liu X, Stamsås GA, Pelliciari S, van Raaphorst R, Dénéréaz J, Kjos M, Murray H, Gruber S, Grossman AD, Veening JW (2021). CcrZ is a spatiotemporal cell cycle regulator that interacts with FtsZ and controls DNA replication by modulating the activity of DnaA. Nat Microbiol. doi.org/10.1038/s41564-021-00949-1. Commentary.
Ovchinnikov K, Kranjec C, Telke A, Kjos M, Thorstensen T, Carlsen H, Scherer S, Diep DB (2021). A strong synergy between the thiopeptide bacteriocin micrococcin P1 and rifampicin against MRSA in a murine skin infection model . Front Immunol. 12: 676534. doi.org/10.3389/fimmu.2021.676534.
40. Straume D, Piechowiak KW, Kjos M, Håvarstein LS (2021) Class A PBPs: it is time to rethink traditional paradigms. Mol Microbiol. doi: https://doi.org/10.1111/mmi.14714
Ruud-Winther A, Kjos M, Herigstad ML, Håvarstein LS, Straume D (2021). EloR interacts with the lytic transglycosylase MltG at midcell in Streptococcus pneumoniae R6. J Bacteriol. doi: 10.1128/JB.00691-20. Preprint: https://www.biorxiv.org/content/10.1101/2020.12.18.423453v1.full
Kranjec C*, Morales-Angeles D*, Torrissen Mårli M, Fernandez L, Garcia P, Kjos M*, Diep DB*. (2021) Staphylococcal Biofilms: Challenges and novel therapeutic perspectives. Antibiotics. 10(2):131. https://doi.org/10.3390/antibiotics10020131
Stamsås GA*, Restelli M*, Ducret A, Freton C, Garcia PS, Håvarstein LS, Straume D, Grangeasse C and Kjos M (2020) A CozE homologue contributes to cell size homeostasis of Streptococcus pneumoniae. mBio. 11(5):e02461-20. https://doi.org/10.1128/mBio.02461-20
Fergestad M, Stamsås GA, Morales-Angeles D, Salehian Z, Wasteson Y, Kjos M (2020) PBP2A provides variable levels of protection towards different β-lactams in Staphylococcus aureus RN4220. Microbiol Open. https://doi.org/10.1002/mbo3.1057
Straume D, Piechowiak KW, Olsen S, Stamsås GA, Berg KH, Kjos M, Heggenhougen MV, Alcorlo M, Hermoso J and Håvarstein LS. Class A PBPs have a distinct and unique role in the construction of the pneumococcal cell wall. Proc Natl Acad Sci U S A. doi.org/10.1073/pnas.1917820117. Preprint available at: https://www.biorxiv.org/content/10.1101/665463v1.
van Raaphorst R, Kjos M, Veening JW (2019) BactMAP: an R package for integrating, analyzing and visualizing bacterial microscopy data. Mol Microbiol. Preprint available at: https://www.biorxiv.org/content/10.1101/728782v1
Myrbråten I, Wiull K, Straume D, Salehian Z, Håvarstein LS, Mathiesen G, Kjos M (2019) CRISPR interference for rapid knockdown of essential cell cycle genes in Lactobacillus plantarum. mSphere. 4(2):e00007-19. Editor's pick.
Ruud Winther A, Kjos M, Stamsås GA, Håvarstein LS, Straume D (2019) Prevention of EloR/KhpA heterodimerization by introduction of site-specific amino acid substitutions renders the essential elongasome protein PBP2b redundant in Streptococcus pneumoniae. Sci Rep. 9:3681.
Kjos M. Transcriptional knockdown in pneumococci using CRISPR interference (2019) In: Streptococcus pneumoniae. Methods and Protocols (Iovino F, eds.), Methods in Molecular Biology Springer Protocols, Germany. 1968:89-98. doi: 10.1007/978-1-4939-9199-0_8.
30. Kjos M. Construction of fluorescent pneumococci for in vivo imaging and labelling of the chromosome (2019) In: Streptococcus pneumoniae. Methods and Protocols (Iovino F, eds.), Methods in Molecular Biology Springer Protocols, Germany. 1968:41-51. doi: 10.1007/978-1-4939-9199-0_4.
Lycus P, Soriano-Laguna M, Kjos M, Richardson D, Gates A, Milligan DA, Frostegård Å, Bergaust L, Bakken LR (2018) A bet-hedging strategy of denitrifying bacteria curtails their release of N2O. Proc Natl Acad Sci U S A. 115(46):11820-11825.
Miller E*, Kjos M*, Abrudan M, Roberts IS, Veening JW, Rozen DE (2018) Crosstalk and eavesdropping among quorum sensing peptide signals that regulate bacteriocin production in Streptococcus pneumoniae. ISME J. 12(10):2363-2375. Preprint available at: http://biorxiv.org/content/early/2016/11/11/087247. Se også: 1.
Stamsås GA*, Myrbråten I*, Straume D, Salehian Z, Veening JW, Håvarstein LS, Kjos M (2018) CozEa and CozEb play overlapping and essential roles in controlling cell division in Staphylococcus aureus. Mol Microbiol. 109(5):615-632. Preprint available at: https://www.biorxiv.org/content/early/2018/05/23/256560. Se også: Cover
Moreno-Gámez S, Sorg RA, Domenech A, Kjos M, Weissing FJ, van Doorn GS, Veening JW. Quorum-sensing integrates environmental cues, cell density and cell history to control bacterial competence. Nat Comm. 8(1):854.
Stamsås GS, Straume D, Ruud Winther A, Kjos M, Frantzen CA, Håvarstein LS (2017) Identification of EloR (Spr1851) as a regulator of cell elongation in Streptococcus pneumoniae. Mol Microbiol. 10.1111/mmi.13748
van Raaphorst R*, Kjos M*, Veening JW (2017) Chromosome segregation drives division site selection in Streptococcus pneumoniae. Proc Natl Acad Sci U S A. 114(29):E5959-E5968. *joint first authors. Se også: 1,2.
Liu X, Gallay C, Kjos M, Domenech A, Slager J, van Kessel S, Knoops K, Sorg RA, Zhang JR, Veening JW (2017) High-throughput CRISPRi phenotyping in Streptococcus pneumoniae identifies new essential genes involved in cell wall synthesis and competence development. Mol Syst Biol. 13:931.
Oppegård C, Kjos M, Veening JW, Nissen-Meyer J, Kristensen T (2016) A putative amino acid transporter determines sensitivity to the two-peptide bacteriocin plantaricin JK. MicrobiologyOpen. doi: 10.1002/mbo3.36.
Kjos M*, Miller E*, Slager J, Lake F, Gericke O, Roberts IS, Rozen DE, Veening JW (2016) Antibiotic-induced expression of pneumococcal bacteriocins via regulatory interplay with the competence system. PLoS Pathogens. 12(2):e1005422. *joint first authors.
Nourikyan J*, Kjos M*, Cluzel C, Morlot C, Mercy C, Noirot-Gros MF, Lavergne JP, Guiral S, Veening JW, Grangeasse C. (2015) Autophosphorylation of the bacterial tyrosine kinase CpsD coordinates capsule synthesis and cell division of Streptococcus pneumoniae. PLoS Genetics. 11(9):e1005518. *joint first authors.
Beilharz K, van Raaphorst R, Kjos M, Veening JW (2015) Red fluorescent proteins for gene expression and protein localization studies in Streptococcus pneumoniae and efficient transformation with DNA assembled via the Gibson assembly method. Appl Environ Microbiol. 81(20):7244-52.
Attaiech L, Minnen A, Kjos M, Gruber S, Veening JW (2015) The ParB-parS chromosome segregation system modulates natural competence development in Streptococcus pneumoniae. mBio. 6(4):e00662-15.
Paixão L, Oliveira J, Verissímo A, Vinga S, Lourenço E, Ventura R, Kjos M, Veening JW, Fernandes VE, Andrew PE, Yesilkaya H and Neves AR (2015) Host glycan sugar specific pathways in Streptococcus pneumoniae: galactose as a key sugar in colonisation and infection. PLoS One. 10(3):e0121042.
16. Kjos M, Aprianto R, Fernandes VE, Andrew PW, van Strijp JAG, Nijland R, Veening JW (2015) Bright fluorescent Streptococcus pneumoniae for live cell imaging of host-pathogen interactions. J Bacteriol. 197:807-18.
15. Kjos M*, Oppegård C*, Diep DB, Nes IF, Veening JW, Nissen-Meyer J, Kristiansen T (2014) Sensitivity to the two-peptide bacteriocin lactococcin G is dependent on an enzyme involved in cell-wall synthesis. Mol Microbiol. 92(6):1177-87. *joint first authors.
Slager J, Kjos M, Attaiech L, Veening JW (2014) Antibiotic-induced increase of origin proximal gene copy number triggers bacterial competence. Cell. 157(2):395-406.
Kjos M, Veening JW (2014) Tracking of chromosome dynamics in live Streptococcus pneumoniae reveals that transcription promotes chromosome segregation. Mol Microbiol. 91(6):1088-1105.
Hassan M, Kjos M, Nes IF, Diep DB, Lotfipour F (2014) Antimicrobial peptides from prokaryotes. In: Novel Antimicrobial Agents and Strategies (Phoenix DA, Harris F, Dennison SR, eds.). Wiley-VCH, Germany.
Pinho MG, Kjos M, Veening JW (2013) How to get (a)round: Mechanisms controlling growth and division of coccoid bacteria. Nat Rev Microbiol. 11:601-14.
Hassan M, Kjos M, Nes IF, Diep DB, Lotfipour F (2012) Natural antimicrobial peptides from bacteria: characteristics and potential applications to fight against antibiotic resistance. J Appl Microbiol. 113(4):723-36.
Nes IF, Kjos M, Diep DB (2011) Antimicrobial components of lactic acid bacteria. In: Lactic acid bacteria: microbiological and functional aspects, fourth edition (Lahtinen S, Ouwehand AC, Salminen S, von Wright A, eds.), CRC Press Taylor & Francis, USA.
Kjos M, Borrero J, Opsata M, Birri DJ, Holo H, Cintas LM, Snipen L, Hernandez PE, Nes IF, Diep DB (2011) Target recognition, resistance, immunity and genome mining of class II bacteriocins from Gram-positive bacteria. Microbiology. 157:3256-67.
Kjos M, Nes IF, Diep DB (2011) Mechanisms of resistance to bacteriocins targeting the mannose phosphotransferase system. Appl Environ Microbiol. 77(10):3335-42.
Kjos M, Salehian Z, Nes IF, Diep DB (2010) An extracellular loop of the mannose phosphotransferase system component IIC is responsible for specific targeting by class IIa bacteriocins. J Bacteriol. 192(22):5906-13.
5. Kjos M, Snipen LS, Salehian Z, Nes IF, Diep DB (2010) The Abi proteins and their involvement in bacteriocin self-immunity. J Bacteriol. 192(8):2068-76.
Kjos M, Nes IF, Diep DB (2009) Class II one-peptide bacteriocins target a phylogenetically defined subgroup of mannose phosphotransferase systems on sensitive cells. Microbiology. 155(9):2949-61.
Diep DB, Straume D, Kjos M, Torres C, Nes IF (2009) An overview of the mosaic bacteriocin pln loci from Lactobacillus plantarum. Peptides 30 (8): 1562-74.
Kjos M, Straume D, Nes IF, Diep DB (2009) Transposition of IS10R in Lactococcus lactis. J Appl Microbiol 106(1):288-95.
Straume D, Kjos M, Nes IF, Diep DB (2007) Quorum-sensing based bacteriocin production is down-regulated by N-terminally truncated species of gene activators. Mol Genet Gen. 278 (3): 283-93.
Group members