About us
Research
To achieve our goal of developing tools to control microbial nitrogen transformations, we need fundamental research on the ecophysiology and regulatory biology of the microbes managing reactive nitrogen in the biosphere.
Our applied research includes collaborative projects with agriculture-, fertilizer-, mining-, and wastewater-industries.
We have developed robotized incubation systems for lab-scale investigations of aerobic and anaerobic respiratory metabolism, and a field robot for measuring gas fluxes in field experiments. Read more about the robots used in our research further down on the page!
News

17.04.2026
Congratulations Martin!
Our dear colleague and friend, Martin Menestreau, successfully defended his PhD thesis titled "Unraveling Denitrification in Stutzerimonas and Other Related Bacteria: Phenotypic, Diversity, Control Mechanisms, Genetic Tools, and Stress Responses". We thank you for your scientific contributions to the nitrogen cycle, and wish you nothing but the best in your future endeavors!

20.03.2026
Congratulations Sigrid!
We wish to congratulate our amazing colleague Sigrid Trier Kjær on her successful PhD defense. Her thesis was entitled "Trade-offs between soil carbon sequestration and nitrous oxide emissions.". We wish you well on your further academic journey back home in Denmark and will eagerly follow your future work! However, you will be heavily missed within the Nitrogen group.

11.03.2026
Article co-written by Prof. Åsa Frostegård on biodegradable plastics
Professor Åsa Frostegård is a member of the Scientific Comittee for Food and Environment (VKM), an advisory board for The Norwegian Food Safety Authority and The Norwegian Environment Agency. They published a popular science article on biodegradable plastics and the misconceptions of how degradable and what environmental consequences they might harbour.
Environmental rationale
We strive to unravel how microbes regulate oxic and anoxic respiration. Why is this environmental research?
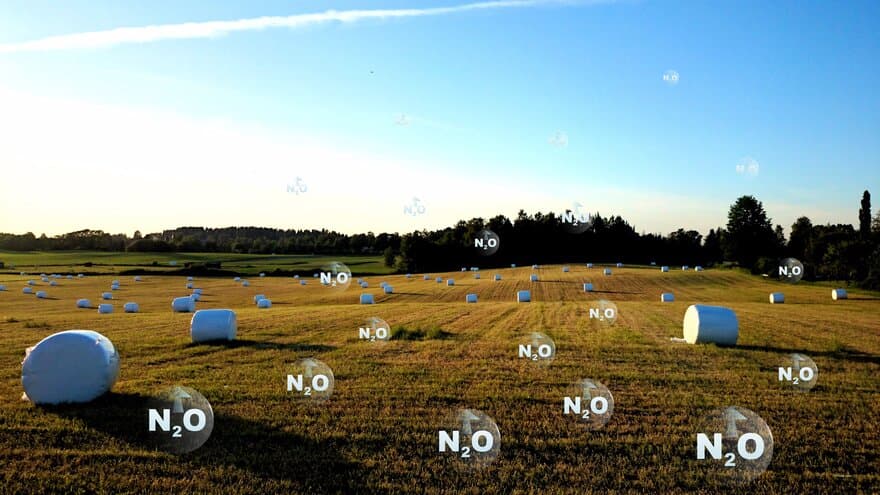
The increasing concentrations of N2O in the atmosphere is due to mankind's interference with the nitrogen cycle.
The ultimate cause is nitrogen enrichment, the proximal N2O sources are nitrogen red/ox reactions by prokaryotes.
We believe that there exist options for reducing N2O emissions, but only if we improve our understanding of the regulatory network at the cellular-, microhabitat-, whole soil-, soil-plant- and ecosystem level. This a goal for NMBU Nitrogen Group.
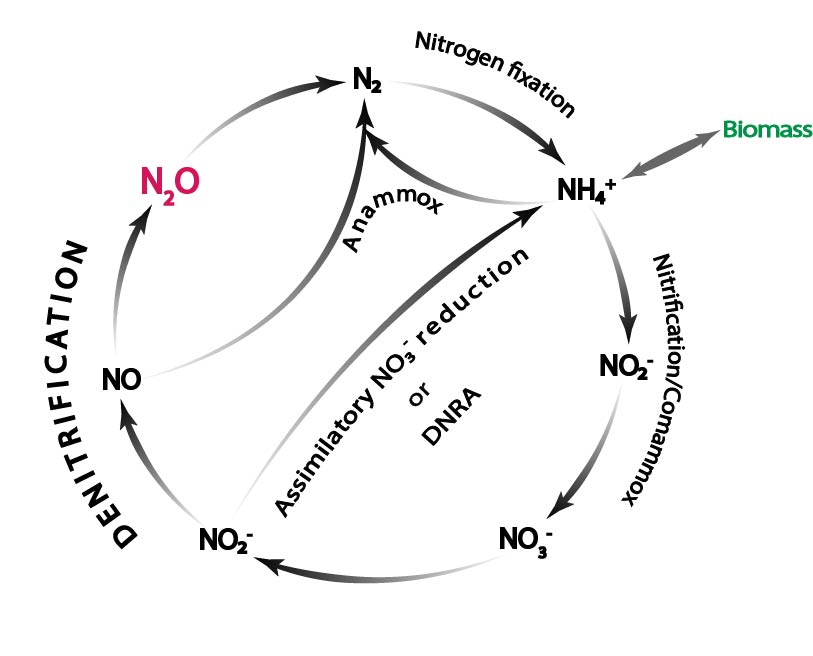
We study the regulatory biology and ecology of denitrifcation, dissimilatory reduction of nitrite to ammonium, and ammonium oxidation.
We use model organisms, such as Paracoccus denitrificans;Nitrosomnas europea, Agrobacterium tumefaciens.
.. not because we believe that they are representative for organisms in nature, but because they are "fast tracks" to new insight, novel phenomena, new concepts.
We are aware that phenomena discovered in model organisms must be validated by experiments with natural populations; which we do.
This is best illustrated by an example:
We discovered that the model organism, Paracoccus denitrificans was unable to make functional N2O reductase at pH 6, although it clearly "tried" by transcribing the gene coding for the enzyme.
Through a series of experiments, we found that the most plausible explanation is that low pH interferes with the assembly of the enzyme in the periplasm, and we hypothesize that Cu-insertion is the critical step. The enzyme, if successfully assembled at pH 7, was perfectly functional at pH<6! This could of course be a peculiarity for Paracoccus denitrificans, but we have provided ample evidence for the opposite by studying natural populations: in all soils tested, we find that the expression of N2O reductase declines with pH, and experiments with bacteria extracted from soil confirmed the peculiar post translatoric phenomenon described for Paracoccus.
We would not need the model strain to discover the effect of pH and the N2O/N2 product ratio of denitrification. The phenomenon was observed more than 60 years ago, but never understood, hence forgotten and rediscovered several times since then. But the experiments with the model strain helped us to understand the cause.
Why is this so intresting? Because the pervasive effect of soil pH on the N2O/N2 product ratio means that we can reduce N2O emissions from soils by liming! Liming increase soil pH, which enhance the expression of N2O reductase, thus lowers the N2O/N2 product ratio of denitrification, hence reduces the N2O emission to the atmosphere!
Environmental research at its best unravels causations.
Anaerobic respiration and N2O production
Our research on denitrification in heterotrophic prokaryotes was initiated in 2005, and our aim was to characterize and understand how they regulate their transition from aerobic to anaerobic respiration in response to oxygen depletion.
At that time, many "nuts and bolts" of the regulatory networks controlling this transition had been unravelled (albeit limited to a few model organisms), but few had attempted to characterize the regulatory phenotypes under stringent experimental conditions.
We developed such routines, and investigated both model organisms and newly isolated organisms. We discovered a range of novel regulatory phenomena and putative regulatory fitness traits such as bet hedging and the formation of persister cells. We also discovered the reason for the apparent inhibition of N2O reductase at low pH. This research is about to change our understanding of denitrifying organisms as sources or sinks for N2O.
A popular hypothesis among microbial ecologists is that denitrifying communities in soils are different regarding their ability to achieve a balanced transition from oxic to anoxic conditions, and that their propensity to emit N2O can be predicted by their genetic "fingerprint". The most naïve version of this hypothesis is that the propensity to emit N2O can be predicted by gene abundance, i.e. that soils which emit much N2O have high nir/nosZ gene abundance and vice versa. Such hypotheses are commonly tested by searching for correlations between gene abundance data and emissions (or the N2O/N2 product ratio of denitrification). No consistent patterns have emerged, however: the numerous correlations found appear to be utterly spurious.
We believe that regulatory biology is a clue to understand the role of denitrifying communities as sources and sinks for N2O in the environment, and that stringent investigations of model organisms can enhance our understanding of how N2O emissions are regulated, hence strengthen our ability to develop mitigations, i.e. actions that reduce the emissions of N2O.
Our investigatins of Paracoccus denitrificans have revealed bet hedging response to oxygen depletion: all cells express nosZ (N2O reductase), while only a minority express NirS. As a result, the population becomes a strong sink for N2O. The phenomen appears to be widespread among denitrifying organisms.
Our investigation of nosZ-expression in P. denitrificans showed that low pH hampers this expression by interering with the maturation of the enzyme in the periplasm. This could be an anecdote for Paracoccus, but this is clearly not the case: All microbial communities tested so far show exactly the same post transcriptional effect of low pH on the making of N2O reductase! We are currently exploring in more detail what goes wrong at low pH. But the results so far suffice to claim that we know why soil pH has a pervasive effect on the N2O/N2 product ratio of denitrification.
A common notion is that N2O reductase is destabilized/destroyed by oxygen. Our experiments with Pa denitrificans show that this is not the case. During aerobic growth, cells with N2O reductase (expressed during anoxic spell) will preserve the enzyme (but diluting it if growing) during aerobic growth. We have also demonstrated that they preserve nitrite reductase during aerobic growth. This explains the "legacy of anoxia" often observed in soil: once exposed to anoxia, the soils responds faster to subsequent anoxic spells.
A long standing notion is that NOR (the enzyme reducing NO to N2O) is a necessity for survival as a denitrifying organism. This is certainly true when grown in pure culture (no organisms can survive for long in the presence of micromolar concentrations of NO). Nevertheless, denitrifying organisms without NOR have been isolated (Falk et al. 2010), and more will be published on this in the near future. Thus, it appears perfectly possible to survive as a denitrifier without NOR, probably because NO toxicity is not a problem under natural conditions (NO scavenged by others). The "Black queen hypothesis" seems relevant here...
Several observations of communities and pure cultures have convinced us that post- transcriptional regulation of electron flow to either O2 or NOx is a clue to understand N2O emissions from soils. Transcription of the denitrification genes appears to be universally repressed by oxygen, but aerobic denitrification is a significant phenomenon, provided that the denitrification proteome has been expressed during foregoing anoxia.
The "post anoxic trauma" is a term used for a community exposed to oxygen after an anoxic spell. The figure above shows that Thauera aminoaromatica is unable to stop denitifying if given a sudden dose of oxygen. The same phenomenon was deomstrated for a denitrifying community by Morley et al 2008 (see publication list). In both cases, nitrous oxide reductase activity was apparently inhibited by oxygen. Our understanding of this phenomenon has deepened over the last years: it appears that the "inhibition" is primarily a "deprivation": that the terminal oxidases compete successfully with N2O reductase for electrons. In contrast, NIR and NOR appear to be more competitive vi a vis terminal oxidases.
Publications
Selected publications from 2020 to 2025 (PDF files are available on request):
2026
Menestreau, M., Milligan, D.A., Sennett, L.B., Bergaust, L.L., Bakken, L.R., Rowley, G., Kjos, M., Shapleigh, J.P., Frostegård, Å.
Distinct Denitrification Phenotypes in Closely Related Bacteria: Clues to Understanding Variations in Nitrite Accumulation Among Stutzerimonas Strains (Environmental Microbiology)
https://doi.org/10.1111/1462-2920.70275
Menestreau, M., Frostegård, Å., Kjos, M.
Strain-specific challenges in applying CRISPR/Cas9-based genome editing in the novel genus Stutzerimonas (Journal of Microbiological Methods)
https://doi.org/10.1016/j.mimet.2026.107478
Maråk, M.M., Duister, I.M., Bakken, L.R., Bergaust, L.L.
Imperatives for anaerobic high cell density culturing by denitrification: technical and physiological perspectives. (Microbial Cell Factories)
2025
Kellermann, R., Kumar, S., Gates, A.J., Bakken, L.R., Spiro, S., Bergaust, L.L.
The Flavohemoglobin Hmp and Nitric Oxide Reductase Restrict Initial nir Expression in the Bet-Hedging Denitrifier Paracoccus denitrificans by Curtailing Hypoxic NO Signalling. (Environmental Microbiology)
2024
Maråk, M.M., Kellermann, R., Bergaust, L.L., Bakken, L.R.
High cell density cultivation by anaerobic respiration. (Microbial Cell Factories)
https://doi.org/10.1186/s12934-024-02595-8
Crowther, T.W., Rappuoli, R., Corinaldesi, C., Danovaro, R., Donohue, T.J., Huisman, J., Stein, L.Y., Timmis, J.K., Timmis, K., Anderson, M.Z., Bakken, L.R., Baylis, M., Behrenfeld, M.J., Boyd, P.W., Brettell, I., Cavicchioli, R., Delavaux, C.S., Foreman, C.M., Jansson, J.K., Koskella, B., Milligan-McClellan, K., North, J.A., Peterson, D., Pizza, M., Ramos, J.L., Reay, D., Remais, J.V., Rich, V.I., Ripple, W.J., Singh, B.K., Smith, G.R., Stewart, F.J., Sullivan, M.B., van den Hoogen, J., van Oppen, M.J.H., Webster, N.S., Zohner, C.M., van Galen, L.G.
Scientists' call to action: Microbes, planetary health, and the Sustainable Development Goals. (Cell)
https://doi.org/10.1016/j.cell.2024.07.051
Sennett, L.B., Roco, C.A., Lim, N.Y.N., Yavitt, J.B., Dörsch, P., Bakken, L.R., Shapleigh, J.P., Frostegård, Å.
Determining how oxygen legacy affects trajectories of soil denitrifier community structure, functional dynamics, and N2O emissions. (Research Square)
https://doi.org/10.21203/rs.3.rs-3837604/v1
Hiis, E.G., Vick, S.H.W., Molstad, L., Røsdal, K., Jonassen, K.R., Winiwarter, W., Bakken, L.R.
Unlocking bacterial potential to reduce farmland N2O emissions. (nature)
2023
Gao, Y., Lycus, P., Arntzen, M.Ø., Shapleigh, J.P., Bakken, LR., Frostegård, Å.
Denitrification by bradyrhizobia under feast and famine and the role of the bc1 complex in securing electrons for N2O reduction. (Applied Environmental Microbiology).
https://doi.org/10.1101/2022.09.29.510233
Grosz, B., Matson, A., Butterbach-Bahl, K., Clough, T., Davidson, E.A., Dechow, R., DelGrosso, S., Diamantopoulos, S., Dörsch, P., Haas, E., He, H., Henri, C.V., Hui, D., Kleineidam, K., Kraus, D., Kuhnert, M., Léonard, J., Mueller, C., Petersen, S.O., Sihi, D., Vogeler, I., Well, R., Yeluripati, J.B., Zhang, J., Scheer, C.
Modeling denitrification: can we report what we don’t know? (Earth and Space Science Open Archive)
10.22541/essoar.168500283.32887682/v1
Song, Y., Wu, D., Ju, X., Dörsch, P., Wang, M., Wang, R., Song, X., Deng, L., Wang, R., Gao, Z., Haider, H., Hou, L., Liu, M., Yu, Y., Yu, H.
Nitrite stimulates HONO and NOx but not N2O emissions in Chinese agricultural soils during nitrification. (Science of the Total Environment)
https://doi.org/10.1016/j.scitotenv.2023.166451
Yu, H., Duan, Y., Mulder, J., Dörsch, P., Zhu, W., Ri, X., Huang, K., Zheng, Z., Kang, R., Wang, C., Quan, Z., Zhu, F., Liu, D., Peng, S., Han, S., Zhang, Y., Fang, Y.
Universal temperature sensitivity of denitrification nitrogen losses in forest soils. (Nature Climate Change)
https://doi.org/10.1038/s41558-023-01708-2
Wei, J., Fontaine, L., Valiente, N., Dörsch, P., Hessen, D.O., Eiler, A.
Trajectories of freshwater microbial genomics and greenhouse gas saturation upon glacial retreat. (Nature Communications)
https://doi.org/10.1038/s41467-023-38806-w
Vekic, T.T., Nadeem, S., Molstad, L., Martinsen, V., Hiis, E.G., Bakken, L.R., Rütting, T., Klemedtsson, L., Dörsch, P.
Effect of calcareous and siliceous amendments on N2O emissions of a grassland soil. (Soil Use and Management)
https://doi.org/10.1111/sum.12913
Jensen, S., Siljanen, H.M.P., Dörsch, P.
Activity and abundance of methanotrophic bacteria in a northern mountainous gradient of wetlands. (Environmental Microbiology Reports)
https://doi.org/10.1111/1758-2229.13137
Highton, M.P., Bakken, L.R., Dörsch, P., Tobias-Hunefeldt, S., Molstad, L., Morales, S.E.
Soil water extract and bacteriome determine N2O emission potential in soils. (Biology and Fertility of Soils)
https://doi.org/10.1007/s00374-022-01690-5
Hiis, E.G., Vick, S.H.W., Molstad, L., Røsdal, K., Jonassen, K.R., Winiwarter, W., Bakken, L.R.
Effective biotechnology for reducing N2O-emissions from farmland: N2O-respiring bacteria vectored by organic waste. (BioRxiv)
https://doi.org/10.1101/2023.10.19.563143
Kellermann, R., Vick, S.H.W., Lindtveit, K., Milligan, D.A., Bergaust, L.L.
Cell density dependent restriction of N2O reductase in denitrifying Pseudomonas spp., a potential driver of N2O emissions in natural systems. (FEMS Microbiology Letters)
https://doi.org/10.1093/femsle/fnad055
Li, H., Song, X., Bakken, L.R., Ju, X.
Reduction of N2O emissions by DMPP depends on the interactions of nitrogen sources (digestate vs. urea) with soil properties. (Journal of Integrative Agriculture)
https://doi.org/10.1016/j.jia.2022.08.009
Lücker, S., Lenferink, W., Bakken, L., Jetten, M., Kessel, M.V.
Hydroxylamine production by Alcaligenes faecalis challenges the paradigm of heterotrophic nitrification. (Research Square, Preprint)
https://doi.org/10.21203/rs.3.rs-3352161/v1
Vick, S.H.W., Jonassen, K.R., Arntzen, M.Ø., Lycus, P., Bakken, L.R.
Meta-omics analyses of dual substrate enrichment culturing of nitrous oxide respiring bacteria suggest that attachment and complex polysaccharide utilisation contributed to the ability of Cloacibacterium strains to reach dominance. (BioRxiv, Preprint)
https://doi.org/10.1101/2023.06.04.543644
Wang, X., Xiang, B., Li, J., Zhang, M., Frostegård, Å., Bakken, L., Zhang, X.
Using adaptive and aggressive N2O-reducing bacteria to augment digestate fertilizer for mitigating N2O emissions from agricultural soils. (Science of the Total Environment)
https://doi.org/10.1016/j.scitotenv.2023.166284
Yoon, S., Heo, H., Han, H., Song, D.U., Bakken, L.R., Frostegård, Å., Yoon, S.
Suggested role of NosZ in preventing N2O inhibition of dissimilatory nitrite reduction to ammonium. (mBio)
2022
Alvarez, C., Andersen, T.O., Eikanger, K.S., Hamfjord, I.W., Niu, P., Weiby, K.V., Årvik, L., Dörsch, P., Hagen, L.H., Pope, P.B., Forberg, D.K., Hustoft H.K., Schwarm, A., Kidane, A.
Methane inhibition by Asparagopsis taxiformis with rumen fluid collected from ventral and central location – a pilot study. (Acta Agriculturae Scandinavica)
https://doi.org/10.1080/09064702.2022.2152196
Allesson, L., Valiente, N., Dörsch, P., Anderson, T., Eiler, A., Hessen, D.O.
Drivers and variability of CO2:O2 saturation along a gradient from boreal to Arctic lakes. (Scientific Reports)
https://doi.org/10.1038/s41598-022-23705-9
Mousavi, H., Cottis, T., Pommeresche, R., Dörsch, P., Solberg, S.Ø.
Plasma-Treated nitrogen-enriched manure does not impose adverse effects on soil fauna feeding activity or springtails and earthworms abundance. (Agronomy)
https://doi.org/10.3390/agronomy12102314
Duffner, C., Kublik, S., Fösel, B., Frostegård, Å., Schloter, M., Bakken, L.R., Schulz, S.
Genotypic and phenotypic characterization of hydrogenotrophic denitrifiers. (Environmental Microbiology)
https://doi.org/10.1111/1462-2920.15921
Frostegård, Å., Vick, S.H., Lim, N.Y., Bakken, L.R., Shapleigh, J.P.
Linking meta-omics to the kinetics of denitrification intermediates reveals pH-dependent causes of N2O emissions and nitrite accumulation in soil. (The ISME Journal)
https://doi.org/10.1038/s41396-021-01045-2
Jonassen, K.R., Hagen, L.H., Vick, S.H., Arntzen, M.Ø., Eijsink, V.G., Frostegård, Å., Lycus, P., Molstad, L., Pope, P.B., Bakken, L.R.
Nitrous oxide respiring bacteria in biogas digestates for reduced agricultural emissions. (The ISME Journal)
https://doi.org/10.1038/s41396-021-01101-x
Jonassen, K.R., Ormåsen, I., Duffner, C., Hvidsten, T.R., Bakken, L.R., Vick, S.H.W.
A Dual Enrichment Strategy Provides Soil- and Digestate-Competent Nitrous Oxide-Respiring Bacteria for Mitigating Climate Forcing in Agriculture. (mBio)
https://doi.org/10.1128/mbio.00788-22
Kellermann, R., Hauge, K., Tjåland, R., Thalmann, S., Bakken, L.R., Bergaust, L.
Preparation for Denitrification and Phenotypic Diversification at the Cusp of Anoxia: a Purpose for N2O Reductase Vis-à-Vis Multiple Roles of O2. (Applied and Environmental Microbiology)
https://doi.org/10.1128/aem.01053-22
Lekberg, Y., Bååth, E., Frostegård, Å., Hammer, E., Hedlund, K., Jansa, J., Kaiser, C., Ramsey, P.W., Řezanka, T., Rousk, J., Wallander, H., Welc, M., Olsson, P.A.
Fatty acid 16:1ω5 as a proxy for arbuscular mycorrhizal fungal biomass: current challenges and ways forward. (Biology and Fertility of Soils)
https://doi.org/10.1007/s00374-022-01670-9
Mousavi, S.A., Gao, Y., Penttinen, P., Frostegård, Å., Paulin, L., Lindström, K.
Using amplicon sequencing of rpoB for identification of inoculant rhizobia from peanut nodules. (Applied Microbiology)
https://doi.org/10.1111/lam.13599
Dong, Y., Yang, J.L., Zhao, X.R., Yang, S.H., Mulder, J., Dörsch, P., Peng, X.H., Zhang, G.L.
Soil acidification and loss of base cations in a subtropical agricultural watershed. (Science of the Total Environment)
https://doi.org/10.1016/j.scitotenv.2022.154338
Munera-Echeverri, J.L., Martinsen, V., Dörsch, P., Obia, A., Mulder, J.
Pigeon pea biochar addition in tropical Arenosol under maize increases gross nitrification rate without an effect on nitrous oxide emission. (Plant and Soil)
https://doi.org/10.1007/s11104-022-05325-4
Dong, Y., Yang, J.L., Zhao, X.R., Yang, S.H., Mulder, J., Dörsch, P., Peng, X.H., Zhang, G.L.
Seasonal dynamics of soil pH and N transformation as affected by N fertilization in subtropical China: An in situ 15N labeling study. (Science of the Total Environment)
https://doi.org/10.1016/j.scitotenv.2021.151596
Dong, Y., Yang, J.L., Zhao, X.R., Yang, S.H., Mulder, J., Dörsch, P., Zhang, G.L.
Nitrate runoff loss and source apportionment in a typical subtropical agricultural watershed. (Environmental Science and Pollution Research)
https://doi.org/10.1007/s11356-021-16935-3
Dong, Y., Yang, J.L., Zhao, X.R., Yang, S.H., Mulder, J., Dörsch, P., Zhang, G.L.
Nitrate leaching and N accumulation in a typical subtropical red soil with N fertilization. (Geoderma)
https://doi.org/10.1016/j.geoderma.2021.115559
Kang, R., Behrendt, T., Mulder, J., Dörsch, P.
Soil Moisture Control of NO Turnover and N2O Release in Nitrogen-Saturated Subtropical Forest Soils. (Forests)
https://doi.org/10.3390/f13081291
Biasi, C., Jokinen, S., Prommer, J., Ambus, P., Dörsch, P., Yu, L., Halas, S., Granger, S., Boeckx, P., Van Nieuland, K., Brüggemann, N., Wissel, H., Voropaev, A., Zilberman, T., Jäntti, H., Trubnikova, T., Welti, N., Voigt, C., Gebus, B., Czupyt, Z., Wanek, W.
Challenges in measuring δ15N in inorganic nitrogen forms: An interlaboratory comparison of three common measurement approaches. (Rapid Communications in Mass Spectrometry)
https://doi.org/10.1002/rcm.9370
Valiente, N., Eiler, A., Allesson, L., Andersen, T., Clayer, F., Crapart, C., Dörsch, P., Fontaine, L., Heuschele, J., Vogt, R.D., Wei, J., de Wit, H.A., Hessen, D.O.
Catchment properties as predictors of greenhouse gas concentrations across a gradient of boreal lakes. (Frontiers in Environmental Science)

Back row from the left: Lars Molstad, Lars Bakken, Peter Dörsch, Rannei Tjåland, Niklas Wickander, Odd Erik Varberg, Seb Rishaug, Marte Maråk, Martin Menestreau, Pawel Lycus and Kjell Rune Jonassen.
Front row from the left: Åsa Frostegård, Linda Bergaust, Sigrid Kjær, Gro Stamsås, Else Marie Aasen, Louise Sennett, Alicia Pascual, Ingrid Duister and Elisabeth Hiis.
Robotics
Watch a brief presentation of our research and field robot:
Robotized solutions for field- and laboratory experiments.
In collaboration with ADIGO, we have developed an autonomous vehicle that measures emissions of N2O and CO2 from the soil surface in field plot experiments.
The robot measures N2O with TDL with a frequency of 1 Hz, allowing emissions to be quantified with a closure time of 3 min.
Tha alpha-version is now in operation in our field experiments, where we test the effects of pH adjustments by lime and mafic minerals.
Documentation of the performance of the robot is found here.
A beta version of the robot will be made commercially available by ADIGO, in collaboration with NMBU.
We are inviting research institutions to join in on this project, named TEMBO.
This video shows the robot in action and how it is controlled from the office:
The video also shows some interesting details regarding the "fast box" technique developed.
Robotics for gas kinetics in laboratory experiments
We have constructed a robotized incubation system (Molstad et al 2007) which allows tracking of electron flow to various electron acceptors (O2, NO2, NO and N2O) in 15 parallel cultures. The system can also be used to measure other relevant trace gases (CO2, CH4, NO and N2O) at or below ambient concentrations, and thus serves in situ flux measurements. Our lab is also equipped with tools for stable isotope analyses (13C, 15N, 18O, D/H) of organic materials and gases (N2O, CH4, CO2).
We have designed several more versions of this system (described here), replacing the microGC with traditional GC and an improved incubator/sampling system that allows parallel incubations of 30 stirred batches (+14 without stirring). One of these also includes H2 and H2S detection. Another, similar system will be installed in 2020.
Gas kinetics analysis
Links calibration tutorials
Kincalc Part 1 - Get StartedKincalc Part 2 - Calibration Basics Kincalc Part 3 - ExtensionsKincalc Part 4 - Exploring and CuringGroup members
Master students
Hedda Ance Myrvold - AnaPro
Helene Gundersen - STARVOX
Espen Waagen - NRBOW
International collaboration
Biology and enzymology of oxic/anoxic respiration
Pawel Michal Lycus (Danish Technological Institute, Denmark)
Oliver Einsle (University of Freiburg, Germany)
Jim Shapleigh (Cornell, USA)
David Richardson and Andrew Gates (University of East Anglia, UK)
Rosa María Martínez-Espinosa (University of Alicante, Spain)
Stephen Spiro (University of Texas, Dallas, USA)
Jeff Cole (University of Birmingham, UK)
Rob van Spanning (Vrije Universiteit, Amsterdam, Netherlands)
Mark van Loosdrecht (Technical University of Delft, Netherlands)
Jörg Simon (Technical University of Darmstadt, Germany)
Nitrification:
Jim Prosser (School of Biological Sciences, University of Aberdeen, UK)Modelling
Olaf Ippisch (Institut für Mathematik, TU Clausthal, Germany)
Frank Bruggeman (VU University, Amsterdam, Netherlands)
Benjamín J. Sánchez (Novonesis, Denmark)
Soils and microbial communities
Sergio Morales (Ottago University, New zealand)
Nitrogen fixation
Kristina Lindström and Leena A. Räsänen (University of Helsinki, Finland)Alumni
PhD scholars
Ricarda Kellerman (2019-2023)
Marte Mølsæter Maråk (2021-2025)
Martin Menestreau (2022-2026)
Sigrid Trier Kjær (2022-2026)
Master students
2025
Sindre Bech Akre
Jakob Waage Lien
Daniels Cernecs
Sara Helene Norland
2024
Seb Rishaug Strand
Odd Erik Varberg
2023
Marte W. Lindholm-Dahle
